Bioinformatic approaches reveal metagenomic characterization of soil microbial community (V体育安卓版)
- PMID: 24691166
- PMCID: "VSports最新版本" PMC3972102
- DOI: 10.1371/journal.pone.0093445
Bioinformatic approaches reveal metagenomic characterization of soil microbial community
Abstract
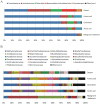
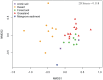
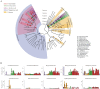
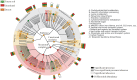
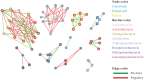
As is well known, soil is a complex ecosystem harboring the most prokaryotic biodiversity on the Earth. In recent years, the advent of high-throughput sequencing techniques has greatly facilitated the progress of soil ecological studies. However, how to effectively understand the underlying biological features of large-scale sequencing data is a new challenge VSports手机版. In the present study, we used 33 publicly available metagenomes from diverse soil sites (i. e. grassland, forest soil, desert, Arctic soil, and mangrove sediment) and integrated some state-of-the-art computational tools to explore the phylogenetic and functional characterizations of the microbial communities in soil. Microbial composition and metabolic potential in soils were comprehensively illustrated at the metagenomic level. A spectrum of metagenomic biomarkers containing 46 taxa and 33 metabolic modules were detected to be significantly differential that could be used as indicators to distinguish at least one of five soil communities. The co-occurrence associations between complex microbial compositions and functions were inferred by network-based approaches. Our results together with the established bioinformatic pipelines should provide a foundation for future research into the relation between soil biodiversity and ecosystem function. .
Conflict of interest statement
"V体育ios版" Figures
References
-
- Daniel R (2005) The metagenomics of soil. Nat Rev Microbiol 3: 470–478. - PubMed
-
- Vogel TM, Simonet P, Jansson JK, Hirsch PR, Tiedje JM, et al. (2009) TerraGenome: a consortium for the sequencing of a soil metagenome. Nat Rev Microbiol 7: 252.
-
- Delmont TO, Robe P, Cecillon S, Clark IM, Constancias F, et al. (2011) Accessing the soil metagenome for studies of microbial diversity. Appl Environ Microbiol 77: 1315–1324. - PMC (VSports app下载) - PubMed
-
- Torsvik V, Øvreås L (2002) Microbial diversity and function in soil: from genes to ecosystems. Curr Opin Microbiol 5: 240–245. - "VSports" PubMed
Publication types
"VSports最新版本" MeSH terms
- Actions (VSports注册入口)
- "VSports最新版本" Actions
"VSports" Substances
VSports - LinkOut - more resources
VSports在线直播 - Full Text Sources
Other Literature Sources
Research Materials (V体育2025版)

